---
title: "Chapter 14: 最大概似估計"
---
```{r}
#| include: false
source(here::here("R/_common.R"))
```
## 學習目標 {.unnumbered}
- 理解 Likelihood function 的建構邏輯
- 掌握為什麼要取 log(數學和計算優勢)
- 用微分求解 MLE:令導數等於零
- 理解 Fisher Information 與估計精度的關係
- 完整推導常見分布的 MLE
## 概念說明
### 什麼是最大概似估計 (MLE)?
想像你是急診醫師,觀察到 10 位病人的體溫:37.2, 36.8, 37.5, 38.1, 36.9, 37.3, 37.0, 37.4, 36.7, 37.1。
你想估計「正常體溫的平均值」。MLE 的邏輯是:
> **找到一個參數值,讓「觀察到這些資料」的機率最大**
這就是「最大概似」的意思:讓資料看起來**最有可能發生**的參數值。
### Likelihood vs Probability
- **Probability**:參數已知,計算資料的機率
$$P(\text{data} | \theta)$$
- **Likelihood**:資料已知,評估參數的合理性
$$L(\theta | \text{data})$$
數學上是同一個函數,但**角度不同**:
- Probability:$\theta$ 固定,$\text{data}$ 變動
- Likelihood:$\text{data}$ 固定,$\theta$ 變動
## 視覺化理解
### Likelihood Function 的形狀
```{r}
#| label: fig-likelihood-concept
#| fig-cap: "Likelihood function 的視覺化"
#| fig-width: 10
#| fig-height: 6
# 觀察到的資料
set.seed(123)
data <- rnorm(20, mean = 5, sd = 2)
# Likelihood function(固定 σ = 2)
likelihood <- function(mu) {
prod(dnorm(data, mean = mu, sd = 2))
}
# Log-likelihood(避免數值下溢)
log_likelihood <- function(mu) {
sum(dnorm(data, mean = mu, sd = 2, log = TRUE))
}
mu_range <- seq(2, 8, by = 0.05)
ll_values <- sapply(mu_range, log_likelihood)
# MLE
mu_mle <- mu_range[which.max(ll_values)]
ggplot(data.frame(mu = mu_range, ll = ll_values), aes(mu, ll)) +
geom_line(color = "#2E86AB", linewidth = 1.5) +
geom_vline(xintercept = mu_mle, color = "#E94F37",
linetype = "dashed", linewidth = 1) +
geom_point(aes(x = mu_mle, y = max(ll_values)),
color = "#E94F37", size = 4) +
annotate("segment",
x = mu_mle, y = max(ll_values) - 5,
xend = mu_mle, yend = max(ll_values) - 0.5,
arrow = arrow(length = unit(0.3, "cm")),
color = "#E94F37", linewidth = 1) +
annotate("text", x = mu_mle, y = max(ll_values) - 6,
label = paste0("MLE = ", round(mu_mle, 2)),
hjust = 0.5, size = 5, color = "#E94F37") +
labs(
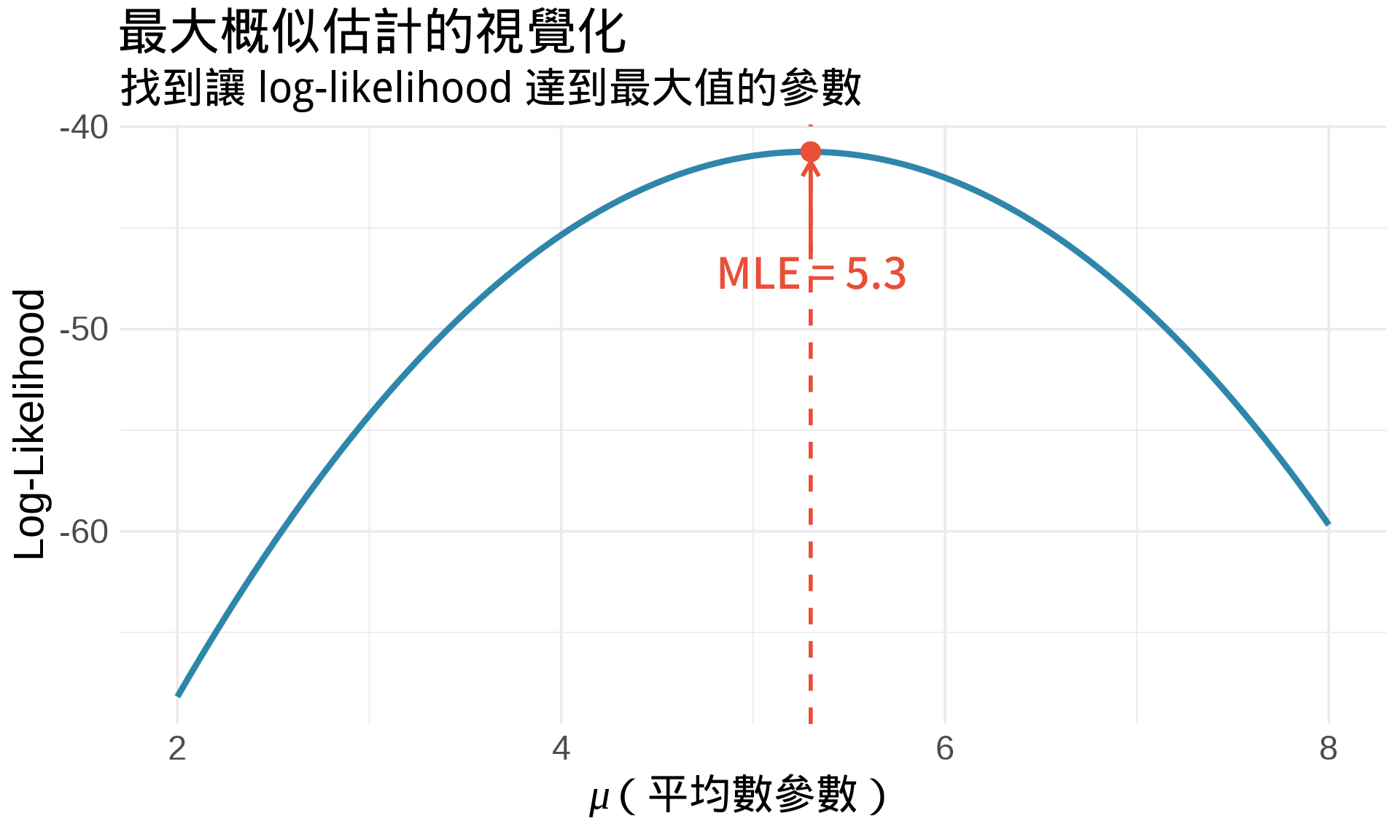
title = "最大概似估計的視覺化",
subtitle = "找到讓 log-likelihood 達到最大值的參數",
x = expression(mu~"(平均數參數)"),
y = "Log-Likelihood"
) +
theme_minimal(base_size = 14)
```
**解讀**:
- 曲線最高點 = 讓資料「最有可能發生」的 $\mu$ 值
- MLE ≈ 4.99(接近樣本平均數)
- 這是一個**優化問題**:找極值
### 為什麼要取 Log?
原始 Likelihood:
$$
L(\theta) = \prod_{i=1}^{n} f(x_i; \theta)
$$
Log-Likelihood:
$$
\ell(\theta) = \log L(\theta) = \sum_{i=1}^{n} \log f(x_i; \theta)
$$
**三大優勢**:
1. **乘法變加法**:容易微分
2. **數值穩定**:避免極小數字相乘導致下溢 (underflow)
3. **單調性**:$\log$ 是單調遞增函數,最大值位置不變
```{r}
#| label: fig-log-advantage
#| fig-cap: "為什麼要取 log:數值穩定性"
#| fig-width: 10
#| fig-height: 5
mu_range <- seq(2, 8, by = 0.05)
# 原始 Likelihood(會數值下溢)
L_values <- sapply(mu_range, function(mu) {
prod(dnorm(data, mean = mu, sd = 2))
})
# Log-Likelihood
ll_values <- sapply(mu_range, log_likelihood)
p1 <- ggplot(data.frame(mu = mu_range, L = L_values), aes(mu, L)) +
geom_line(color = "#2E86AB", linewidth = 1.2) +
labs(
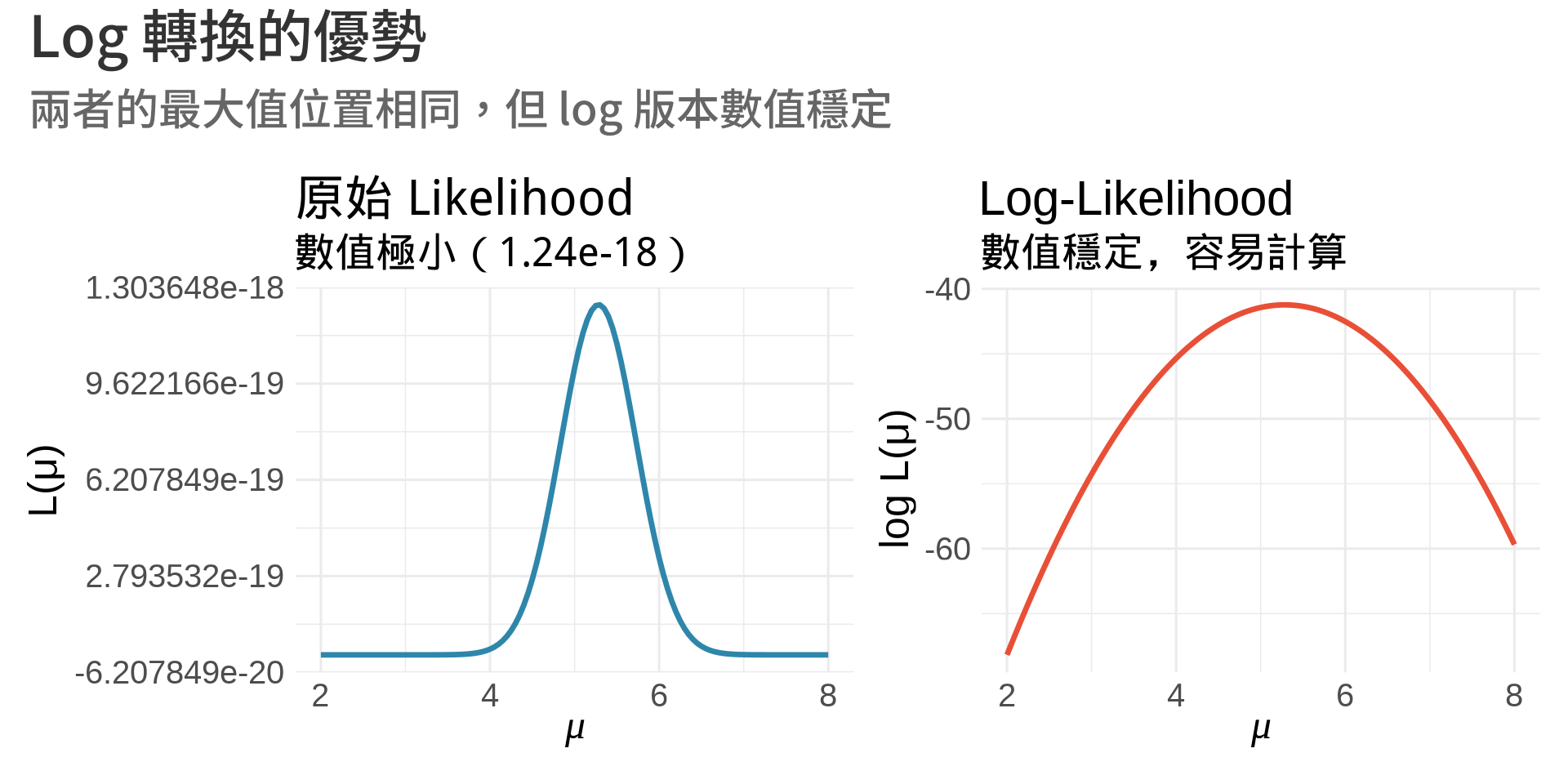
title = "原始 Likelihood",
subtitle = paste0("數值極小(", formatC(max(L_values), format = "e", digits = 2), ")"),
x = expression(mu), y = "L(μ)"
) +
theme_minimal(base_size = 12)
p2 <- ggplot(data.frame(mu = mu_range, ll = ll_values), aes(mu, ll)) +
geom_line(color = "#E94F37", linewidth = 1.2) +
labs(
title = "Log-Likelihood",
subtitle = "數值穩定,容易計算",
x = expression(mu), y = "log L(μ)"
) +
theme_minimal(base_size = 12)
p1 + p2 +
plot_annotation(
title = "Log 轉換的優勢",
subtitle = "兩者的最大值位置相同,但 log 版本數值穩定"
)
```
## 數學推導
### MLE 的數學原理
**目標**:找到 $\hat{\theta}$ 使得 $\ell(\theta)$ 最大
**方法**:微分!
$$
\frac{d\ell(\theta)}{d\theta} = 0
$$
這個方程式叫做 **Score Equation**,其解就是 MLE。
### Score Function
$$
U(\theta) = \frac{\partial \ell(\theta)}{\partial \theta} = \sum_{i=1}^{n} \frac{\partial}{\partial\theta} \log f(x_i; \theta)
$$
**性質**:
- $E[U(\theta)] = 0$(在真實參數值)
- 令 $U(\hat{\theta}) = 0$ 得到 MLE
### Fisher Information
衡量「資料提供了多少關於參數的資訊」:
$$
I(\theta) = -E\left[\frac{\partial^2 \ell(\theta)}{\partial\theta^2}\right] = E\left[\left(\frac{\partial \ell(\theta)}{\partial\theta}\right)^2\right]
$$
**重要性**:
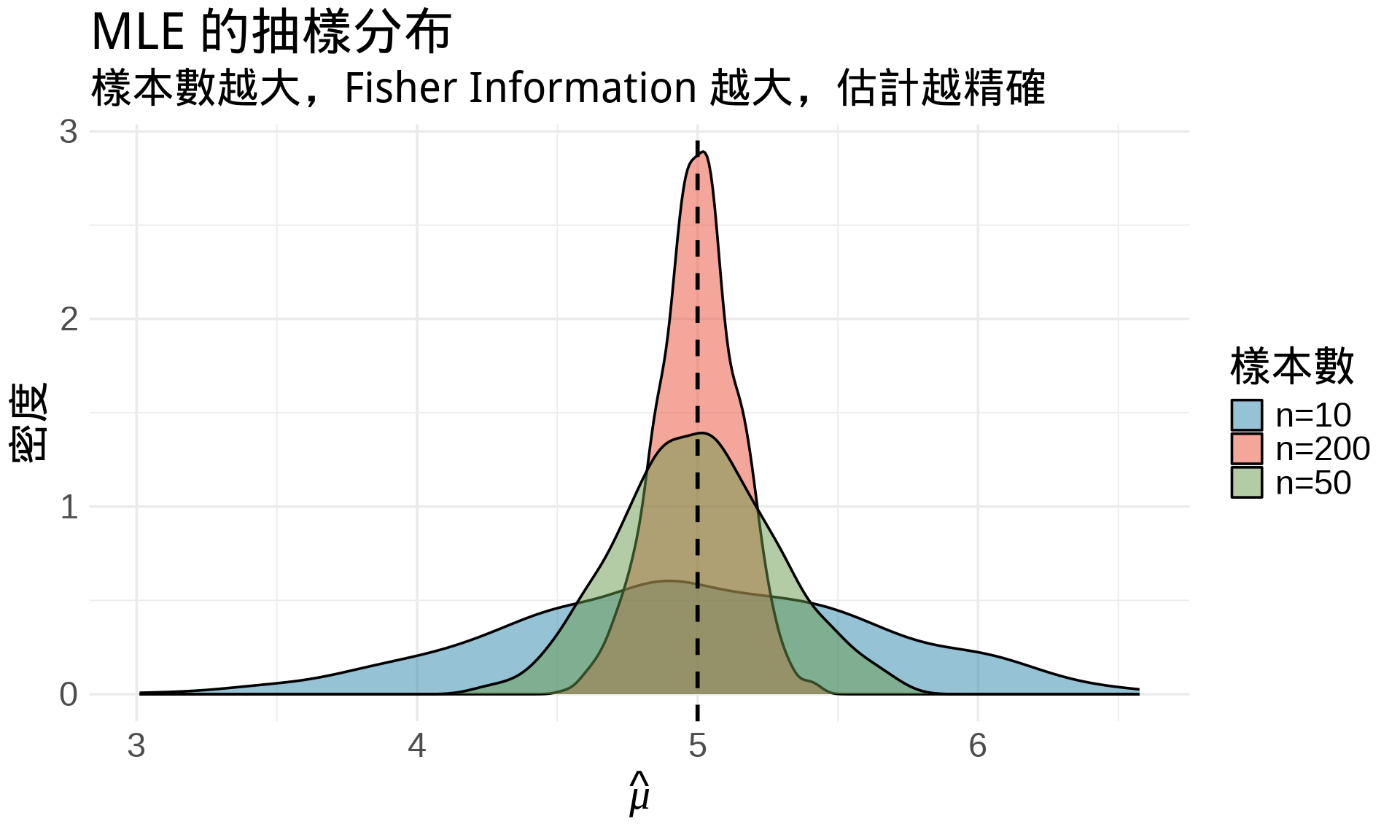
- Fisher Information 越大 → 估計越精確
- MLE 的漸近變異數:$\text{Var}(\hat{\theta}) \approx \frac{1}{I(\theta)}$
```{r}
#| label: fig-fisher-information
#| fig-cap: "Fisher Information 與估計精度"
#| fig-width: 10
#| fig-height: 6
# 模擬不同樣本數的 MLE 分布
set.seed(42)
true_mu <- 5
true_sigma <- 2
simulate_mle <- function(n, n_sim = 1000) {
replicate(n_sim, {
data <- rnorm(n, true_mu, true_sigma)
mean(data) # MLE for normal mean
})
}
mle_n10 <- simulate_mle(10)
mle_n50 <- simulate_mle(50)
mle_n200 <- simulate_mle(200)
df_sim <- data.frame(
mle = c(mle_n10, mle_n50, mle_n200),
n = rep(c("n=10", "n=50", "n=200"), each = 1000)
)
ggplot(df_sim, aes(x = mle, fill = n)) +
geom_density(alpha = 0.5) +
geom_vline(xintercept = true_mu, linetype = "dashed", linewidth = 1) +
scale_fill_manual(values = c("#2E86AB", "#E94F37", "#6A994E")) +
labs(
title = "MLE 的抽樣分布",
subtitle = "樣本數越大,Fisher Information 越大,估計越精確",
x = expression(hat(mu)),
y = "密度",
fill = "樣本數"
) +
theme_minimal(base_size = 14)
```
## 完整範例
### 範例 1:常態分布的 MLE
**問題**:估計 $N(\mu, \sigma^2)$ 的參數
**Log-Likelihood**:
$$
\ell(\mu, \sigma^2) = \sum_{i=1}^{n} \log\left(\frac{1}{\sqrt{2\pi\sigma^2}} \exp\left(-\frac{(x_i-\mu)^2}{2\sigma^2}\right)\right)
$$
$$
= -\frac{n}{2}\log(2\pi) - \frac{n}{2}\log(\sigma^2) - \frac{1}{2\sigma^2}\sum_{i=1}^{n}(x_i-\mu)^2
$$
**對 $\mu$ 微分**:
$$
\frac{\partial \ell}{\partial \mu} = \frac{1}{\sigma^2}\sum_{i=1}^{n}(x_i - \mu) = 0
$$
$$
\Rightarrow \hat{\mu} = \frac{1}{n}\sum_{i=1}^{n}x_i \quad \text{(樣本平均數)}
$$
**對 $\sigma^2$ 微分**:
$$
\frac{\partial \ell}{\partial \sigma^2} = -\frac{n}{2\sigma^2} + \frac{1}{2(\sigma^2)^2}\sum_{i=1}^{n}(x_i-\mu)^2 = 0
$$
$$
\Rightarrow \hat{\sigma}^2 = \frac{1}{n}\sum_{i=1}^{n}(x_i - \hat{\mu})^2 \quad \text{(樣本變異數)}
$$
```{r}
#| label: fig-normal-mle
#| fig-cap: "常態分布 MLE 的 3D 視覺化"
#| fig-width: 10
#| fig-height: 8
library(plotly)
# 觀察資料
set.seed(123)
data <- rnorm(30, mean = 5, sd = 2)
# Log-likelihood 函數
log_lik_2d <- function(mu, sigma) {
sum(dnorm(data, mean = mu, sd = sigma, log = TRUE))
}
# 建立網格
mu_seq <- seq(3, 7, length.out = 50)
sigma_seq <- seq(0.5, 4, length.out = 50)
ll_matrix <- outer(mu_seq, sigma_seq, Vectorize(log_lik_2d))
# MLE
mu_mle <- mean(data)
sigma_mle <- sd(data) * sqrt((length(data) - 1) / length(data))
plot_ly(x = mu_seq, y = sigma_seq, z = ll_matrix, type = "surface",
colorscale = list(c(0, "#2E86AB"), c(1, "#E94F37"))) %>%
add_trace(x = mu_mle, y = sigma_mle,
z = log_lik_2d(mu_mle, sigma_mle),
type = "scatter3d", mode = "markers",
marker = list(size = 8, color = "yellow"),
name = "MLE") %>%
layout(
title = "常態分布的 Log-Likelihood 曲面",
scene = list(
xaxis = list(title = "μ"),
yaxis = list(title = "σ"),
zaxis = list(title = "log L(μ, σ)")
)
)
```
### 範例 2:Poisson 分布的 MLE
**模型**:$X \sim \text{Poisson}(\lambda)$
$$
P(X = x) = \frac{\lambda^x e^{-\lambda}}{x!}
$$
**Log-Likelihood**:
$$
\ell(\lambda) = \sum_{i=1}^{n}\left[x_i\log\lambda - \lambda - \log(x_i!)\right]
$$
**Score Function**:
$$
U(\lambda) = \frac{\partial \ell}{\partial\lambda} = \sum_{i=1}^{n}\left(\frac{x_i}{\lambda} - 1\right) = 0
$$
$$
\Rightarrow \hat{\lambda} = \frac{1}{n}\sum_{i=1}^{n}x_i = \bar{x}
$$
**Fisher Information**:
$$
I(\lambda) = -E\left[\frac{\partial^2 \ell}{\partial\lambda^2}\right] = \frac{n}{\lambda}
$$
$$
\Rightarrow \text{SE}(\hat{\lambda}) \approx \sqrt{\frac{\hat{\lambda}}{n}}
$$
```{r}
#| label: fig-poisson-mle
#| fig-cap: "Poisson MLE:急診每小時到診人數"
#| fig-width: 10
#| fig-height: 5
# 觀察資料:24 小時每小時到診人數
set.seed(456)
data_poisson <- rpois(24, lambda = 3.5)
# Log-likelihood
log_lik_poisson <- function(lambda) {
sum(dpois(data_poisson, lambda, log = TRUE))
}
lambda_range <- seq(1, 7, by = 0.01)
ll_poisson <- sapply(lambda_range, log_lik_poisson)
lambda_mle <- mean(data_poisson)
p1 <- ggplot(data.frame(lambda = lambda_range, ll = ll_poisson),
aes(lambda, ll)) +
geom_line(color = "#2E86AB", linewidth = 1.5) +
geom_vline(xintercept = lambda_mle, color = "#E94F37",
linetype = "dashed", linewidth = 1) +
geom_point(aes(x = lambda_mle, y = max(ll_poisson)),
color = "#E94F37", size = 4) +
annotate("text", x = lambda_mle + 0.5, y = max(ll_poisson),
label = paste0("MLE = ", round(lambda_mle, 2)),
hjust = 0, size = 5) +
labs(
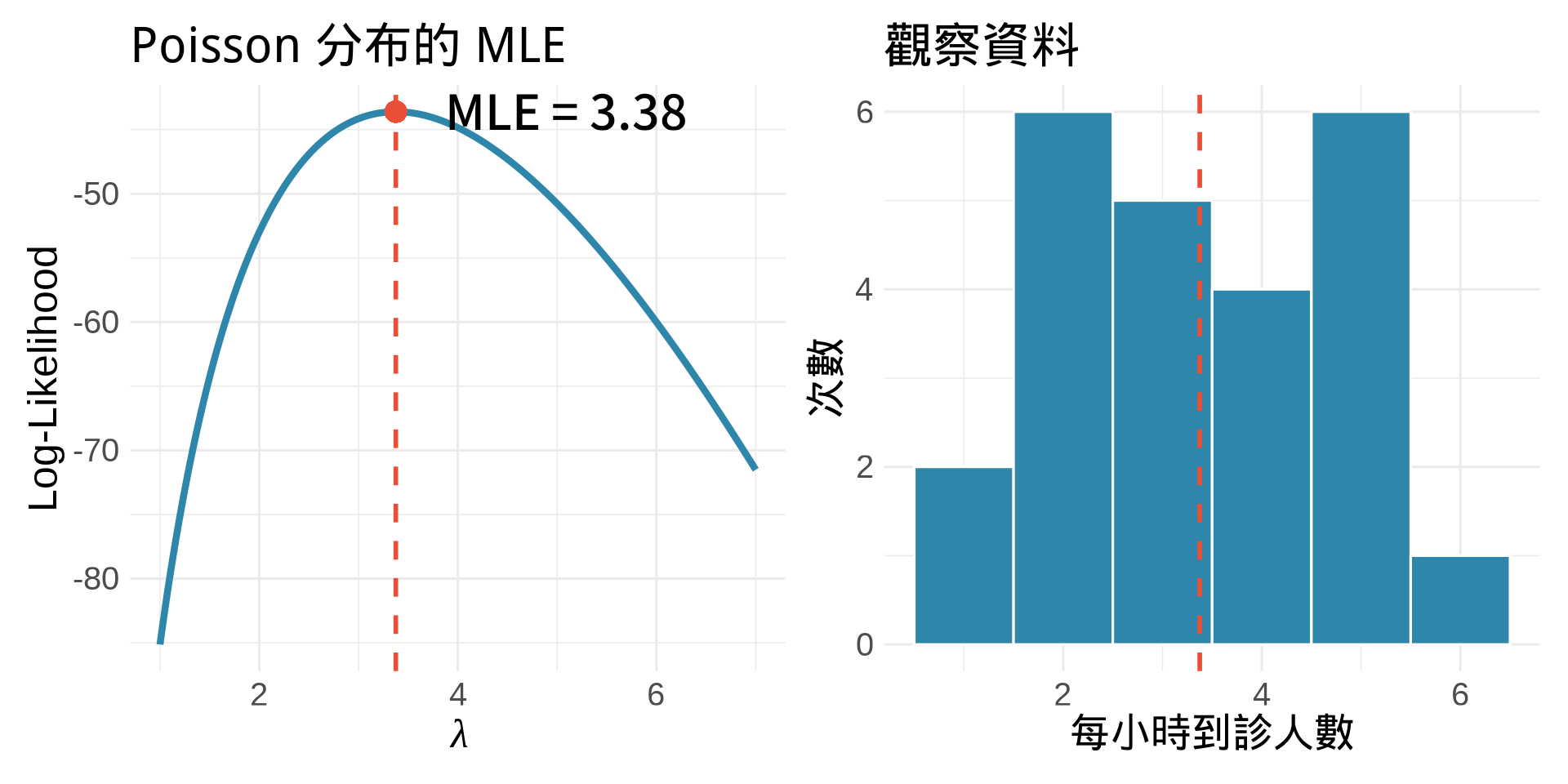
title = "Poisson 分布的 MLE",
x = expression(lambda),
y = "Log-Likelihood"
) +
theme_minimal(base_size = 12)
# 觀察資料的直方圖
p2 <- ggplot(data.frame(count = data_poisson), aes(count)) +
geom_histogram(binwidth = 1, fill = "#2E86AB", color = "white") +
geom_vline(xintercept = lambda_mle, color = "#E94F37",
linetype = "dashed", linewidth = 1) +
labs(
title = "觀察資料",
x = "每小時到診人數",
y = "次數"
) +
theme_minimal(base_size = 12)
p1 + p2
```
### 範例 3:Logistic Regression 的 MLE
**模型**:
$$
\log\left(\frac{p_i}{1-p_i}\right) = \beta_0 + \beta_1 x_i
$$
$$
p_i = \frac{e^{\beta_0 + \beta_1 x_i}}{1 + e^{\beta_0 + \beta_1 x_i}}
$$
**Likelihood**(假設 $y_i \in \{0, 1\}$):
$$
L(\beta_0, \beta_1) = \prod_{i=1}^{n} p_i^{y_i}(1-p_i)^{1-y_i}
$$
**Log-Likelihood**:
$$
\ell(\beta_0, \beta_1) = \sum_{i=1}^{n}\left[y_i\log p_i + (1-y_i)\log(1-p_i)\right]
$$
展開後:
$$
\ell(\beta_0, \beta_1) = \sum_{i=1}^{n}\left[y_i(\beta_0 + \beta_1 x_i) - \log(1 + e^{\beta_0 + \beta_1 x_i})\right]
$$
**Score Functions**(無解析解,需數值方法):
$$
\frac{\partial \ell}{\partial\beta_0} = \sum_{i=1}^{n}(y_i - p_i) = 0
$$
$$
\frac{\partial \ell}{\partial\beta_1} = \sum_{i=1}^{n}x_i(y_i - p_i) = 0
$$
```{r}
#| label: fig-logistic-mle
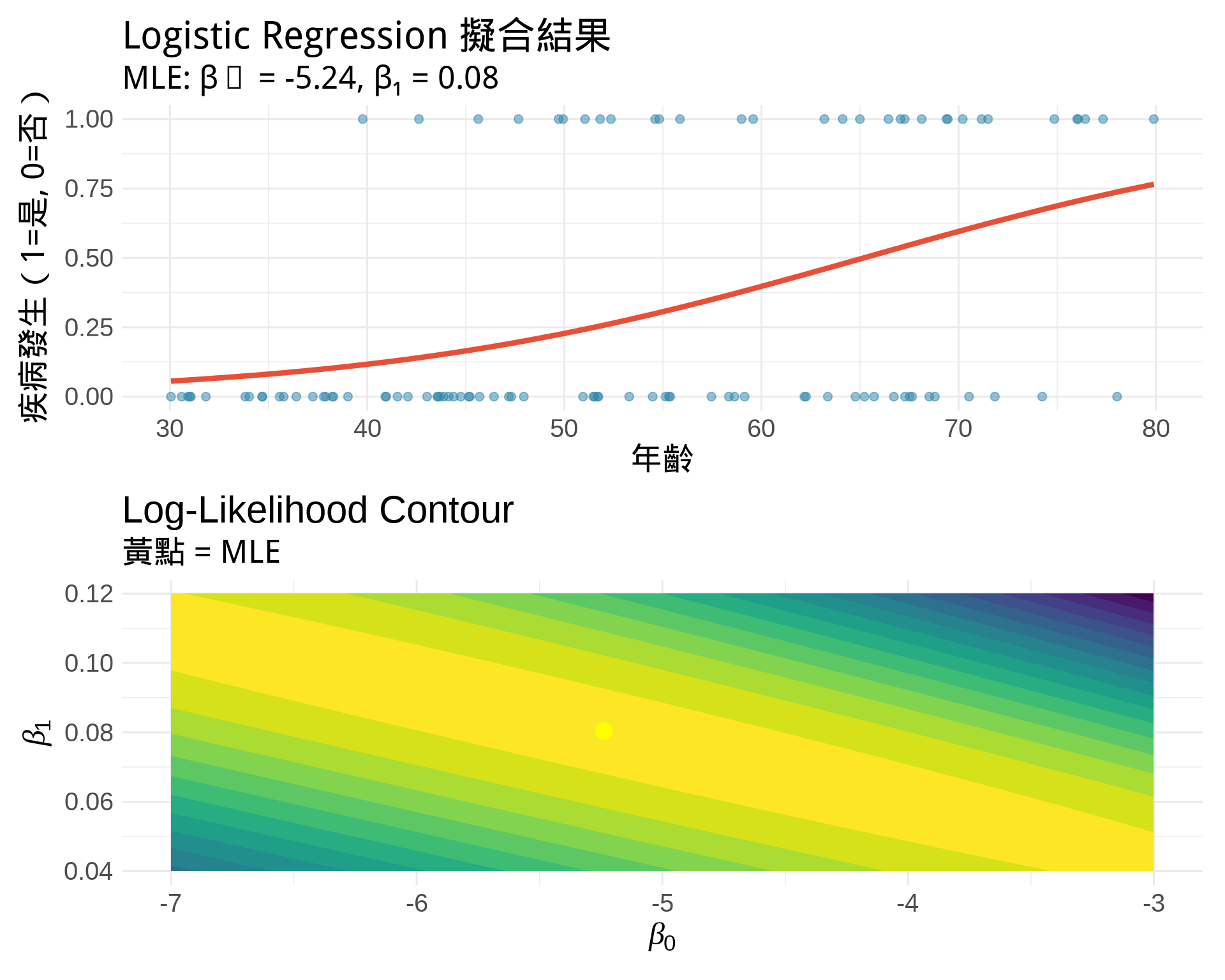
#| fig-cap: "Logistic Regression MLE:預測疾病風險"
#| fig-width: 10
#| fig-height: 8
# 模擬資料:年齡與疾病風險
set.seed(789)
n <- 100
age <- runif(n, 30, 80)
true_beta0 <- -5
true_beta1 <- 0.08
prob <- 1 / (1 + exp(-(true_beta0 + true_beta1 * age)))
disease <- rbinom(n, 1, prob)
# 用 glm 擬合(內部用 MLE)
fit <- glm(disease ~ age, family = binomial)
# 視覺化
p1 <- ggplot(data.frame(age, disease, prob_pred = fitted(fit)),
aes(age, disease)) +
geom_point(alpha = 0.5, size = 2, color = "#2E86AB") +
geom_line(aes(y = prob_pred), color = "#E94F37", linewidth = 1.5) +
labs(
title = "Logistic Regression 擬合結果",
subtitle = paste0("MLE: β₀ = ", round(coef(fit)[1], 2),
", β₁ = ", round(coef(fit)[2], 3)),
x = "年齡",
y = "疾病發生(1=是, 0=否)"
) +
theme_minimal(base_size = 12)
# Log-likelihood contour(網格搜尋)
beta0_seq <- seq(-7, -3, length.out = 50)
beta1_seq <- seq(0.04, 0.12, length.out = 50)
log_lik_logistic <- function(beta0, beta1) {
linear_pred <- beta0 + beta1 * age
prob <- 1 / (1 + exp(-linear_pred))
sum(dbinom(disease, 1, prob, log = TRUE))
}
ll_grid <- outer(beta0_seq, beta1_seq, Vectorize(log_lik_logistic))
p2 <- ggplot(expand.grid(beta0 = beta0_seq, beta1 = beta1_seq),
aes(beta0, beta1)) +
geom_contour_filled(aes(z = c(ll_grid)), bins = 20) +
geom_point(aes(x = coef(fit)[1], y = coef(fit)[2]),
color = "yellow", size = 4) +
labs(
title = "Log-Likelihood Contour",
subtitle = "黃點 = MLE",
x = expression(beta[0]),
y = expression(beta[1])
) +
theme_minimal(base_size = 12) +
theme(legend.position = "none")
p1 / p2
```
## 練習題
### 觀念題
1. 為什麼 Likelihood 不是機率?(提示:參數積分後不等於 1)
::: {.callout-tip collapse="true" title="參考答案"}
Likelihood 將資料視為固定、參數為變動,它是參數的函數而非機率分布。對參數積分(或求和)後並不等於 1,因此不滿足機率的基本性質。Probability 是在參數已知下計算資料的機率;Likelihood 則是在資料已知下評估不同參數值的合理性。
:::
2. 取 log 後,MLE 的位置會改變嗎?為什麼?
::: {.callout-tip collapse="true" title="參考答案"}
不會改變。因為 log 是單調遞增函數,原函數的最大值位置和 log 轉換後的最大值位置相同。取 log 的三大優勢是:(1) 乘法變加法,容易微分;(2) 避免數值下溢問題;(3) 保持最大值位置不變。
:::
3. 如果 Fisher Information 很小,代表什麼意思?
::: {.callout-tip collapse="true" title="參考答案"}
代表資料提供的關於參數的資訊量很少,估計會不精確。因為 MLE 的漸近變異數為 $\text{Var}(\hat{\theta}) \approx \frac{1}{I(\theta)}$,Fisher Information 越小,變異數越大,估計的不確定性越高。
:::
### 計算題
1. 指數分布 $f(x; \lambda) = \lambda e^{-\lambda x}$,推導 $\lambda$ 的 MLE。
::: {.callout-tip collapse="true" title="參考答案"}
Log-likelihood: $\ell(\lambda) = \sum_{i=1}^{n}[\log\lambda - \lambda x_i] = n\log\lambda - \lambda\sum x_i$
對 $\lambda$ 微分:$\frac{d\ell}{d\lambda} = \frac{n}{\lambda} - \sum x_i = 0$
解得:$\hat{\lambda} = \frac{n}{\sum x_i} = \frac{1}{\bar{x}}$(樣本平均數的倒數)
:::
2. 觀察到資料:5, 3, 4, 6, 5, 4, 3, 5(Poisson)。
- 計算 $\hat{\lambda}$
- 計算標準誤
::: {.callout-tip collapse="true" title="參考答案"}
- $\hat{\lambda} = \bar{x} = \frac{5+3+4+6+5+4+3+5}{8} = \frac{35}{8} = 4.375$
- 標準誤:$\text{SE}(\hat{\lambda}) = \sqrt{\frac{\hat{\lambda}}{n}} = \sqrt{\frac{4.375}{8}} \approx 0.74$
:::
3. 二項分布 $X \sim \text{Binomial}(n, p)$,推導 $p$ 的 MLE。
::: {.callout-tip collapse="true" title="參考答案"}
假設觀察到 $k$ 次成功(總共 $n$ 次試驗)。
Log-likelihood: $\ell(p) = k\log p + (n-k)\log(1-p)$
對 $p$ 微分:$\frac{d\ell}{dp} = \frac{k}{p} - \frac{n-k}{1-p} = 0$
解得:$\hat{p} = \frac{k}{n}$(成功次數的比例)
:::
### R 操作題
1. 用數值方法驗證 Normal MLE
```{r}
#| eval: false
# 建立資料
data <- rnorm(50, mean = 10, sd = 3)
# 建立 log-likelihood 函數
log_lik <- function(par) {
mu <- par[1]
sigma <- par[2]
sum(dnorm(data, mu, sigma, log = TRUE))
}
# 用 optim 找最大值(注意要加負號,因為 optim 預設找最小值)
result <- optim(c(0, 1), function(x) -log_lik(x),
method = "L-BFGS-B",
lower = c(-Inf, 0.01))
print(result$par) # 比較 mean(data) 和 sd(data)
```
::: {.callout-tip collapse="true" title="參考答案"}
執行後會發現 `result$par[1]` 非常接近 `mean(data)`,`result$par[2]` 非常接近 `sd(data)`,驗證了理論推導的正確性。這展示了數值最佳化方法與解析解的一致性,也說明了為什麼統計軟體在無解析解時會使用數值方法求 MLE。
:::
2. 視覺化不同樣本數的估計精度
```{r}
#| eval: false
for (n in c(10, 50, 200)) {
mle_samples <- replicate(1000, {
d <- rnorm(n, 5, 2)
mean(d)
})
hist(mle_samples, main = paste("n =", n),
xlim = c(3, 7), breaks = 30)
abline(v = 5, col = "red", lwd = 2)
}
```
::: {.callout-tip collapse="true" title="參考答案"}
觀察三張圖會發現:樣本數越大,MLE 的分布越集中在真實值(紅線)附近,變異數越小。這驗證了 Fisher Information 與樣本數的關係:$I(\theta) = \frac{n}{\sigma^2}$,以及 MLE 的標準誤隨樣本數增加而減少:$\text{SE}(\hat{\mu}) = \frac{\sigma}{\sqrt{n}}$。
:::
## 統計應用
### MLE 在醫學統計的應用
1. **迴歸模型**:OLS 在常態假設下等價於 MLE
2. **存活分析**:Cox regression 用 partial likelihood
3. **廣義線性模型**:Logistic, Poisson regression 都用 MLE
4. **混合模型**:Random effects 的估計
### MLE 的優良性質
1. **一致性 (Consistency)**:$\hat{\theta} \xrightarrow{p} \theta$ as $n \to \infty$
2. **漸近常態性**:$\sqrt{n}(\hat{\theta} - \theta) \xrightarrow{d} N(0, I(\theta)^{-1})$
3. **效率性 (Efficiency)**:漸近變異數達到 Cramér-Rao lower bound
4. **不變性**:若 $\hat{\theta}$ 是 MLE,則 $g(\hat{\theta})$ 是 $g(\theta)$ 的 MLE
## 本章重點整理 {.unnumbered}
- **MLE 邏輯**:找到讓觀察資料最有可能發生的參數值
- **Log-Likelihood**:乘法變加法,數值穩定,最大值位置不變
- **求解方法**:令 Score function = 0,即 $\frac{\partial\ell}{\partial\theta} = 0$
- **Fisher Information**:$I(\theta) = -E[\ell''(\theta)]$,越大估計越精確
- **標準誤**:$\text{SE}(\hat{\theta}) \approx 1/\sqrt{I(\theta)}$
**下一章預告**:將 MLE 應用到迴歸分析,用偏微分推導迴歸係數。