flowchart LR
A[Baseline] -->|Immortal time| B[Treatment received]
B -->|Follow-up| C[Event or censoring]
A -->|Naive analysis credits<br/>all time to treatment| C
7 Time-Varying Exposures
When treatment or exposure status changes during follow-up, standard Cox regression with a baseline-only covariate introduces immortal time bias. This module implements landmark analysis and time-dependent Cox models to handle this correctly.
7.1 Immortal Time Bias
If treatment is received at some point after baseline, the time between baseline and treatment receipt is “immortal” — the patient must survive to receive treatment. Coding treatment as a fixed baseline variable credits this survival time to the treatment group, biasing the hazard ratio downward1.
7.2 Script Pipeline
| Script | Purpose |
|---|---|
01_landmark_analysis.R |
Landmark analysis at fixed time points |
02_tdc_data_prep.R |
Prepare counting-process data with tmerge |
03_tdc_cox.R |
Time-dependent Cox model |
04_immortal_time_check.R |
Compare naive vs corrected estimates |
05_visualization.R |
Forest plot of naive vs corrected HR |
7.3 Landmark Analysis
Landmark analysis restricts the cohort to patients alive and event-free at a fixed landmark time, classifying treatment status as of that moment:
landmark_time <- 90 # days
df_landmark <- df |>
filter(time > landmark_time) |>
mutate(
treatment = ifelse(treatment_date <= landmark_time, 1, 0),
time_from_landmark = time - landmark_time
)
coxph(Surv(time_from_landmark, status) ~ treatment + age, data = df_landmark)The landmark time should be clinically motivated (e.g., 90 days post-diagnosis for transplant studies). Sensitivity analyses at multiple landmarks strengthen the finding.
7.4 Time-Dependent Cox with tmerge
For a more flexible approach, tmerge creates counting-process intervals that update exposure status at the exact time of change:
library(survival)
df_td <- tmerge(df, df, id = id, tstop = time) |>
tmerge(treatment_df, id = id, trt = tdc(treatment_date))
td_fit <- coxph(Surv(tstart, tstop, status) ~ trt + age, data = df_td)
summary(td_fit)7.5 Naive vs Corrected Comparison
Script 04_immortal_time_check.R fits both the naive (fixed baseline) and corrected (time-dependent) models side by side:
Naive HR: 0.62 (0.48-0.80) # biased downward
Corrected HR: 0.85 (0.65-1.11) # unbiasedShowing the naive and corrected estimates together makes the magnitude of immortal time bias explicit, which strengthens the manuscript’s methodological rigour.
7.6 Running the Module
make analyze-time-varying PROJECT=my-study7.7 Demo: Immortal Time Bias (Scenario 5)
N=800, treatment switch at random time for 33.6% of patients (269/800). True HR = 0.80.
7.7.1 The Bias Revealed
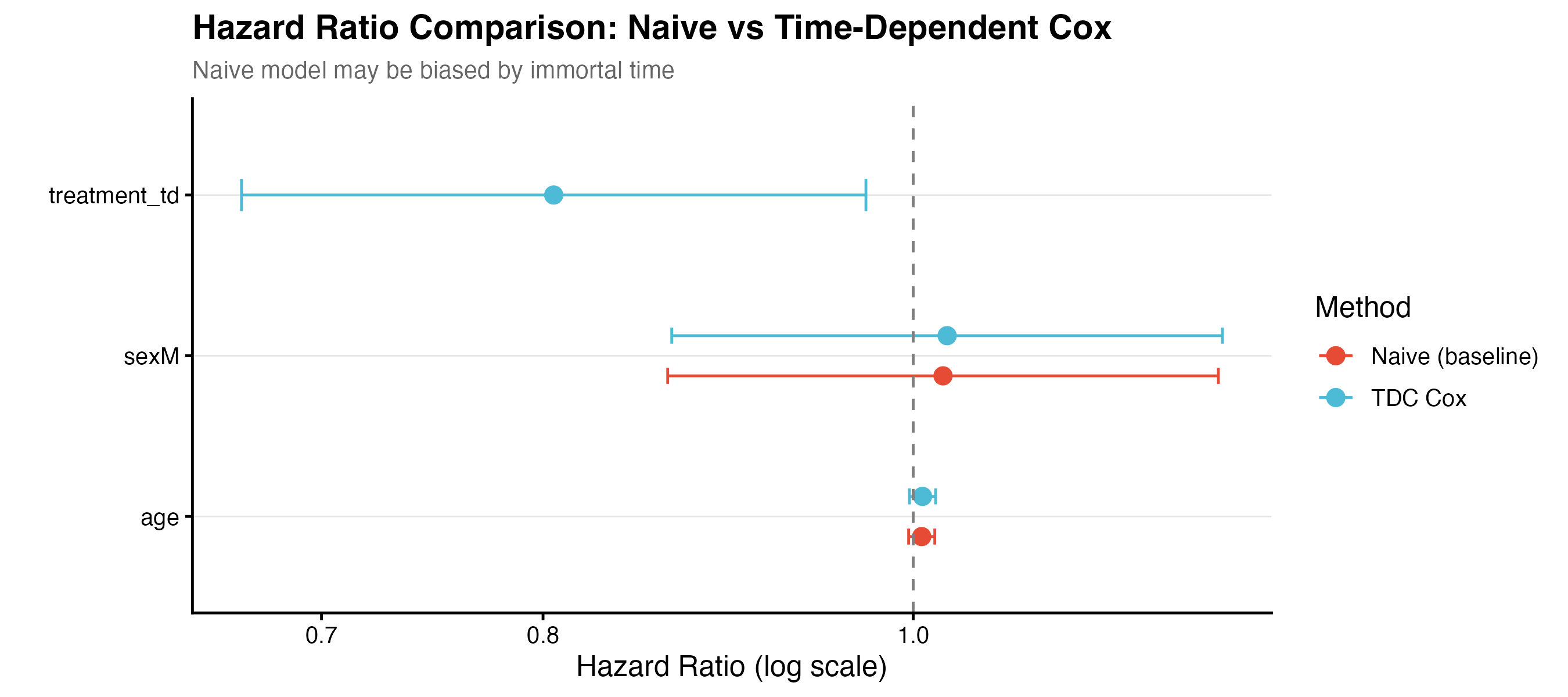
The naive analysis assigned treatment as a fixed baseline variable. Because all patients start untreated and only some switch later, the naive model cannot estimate a treatment HR (the baseline treatment coefficient is NA). The time-dependent Cox model correctly handles the switching and recovers HR = 0.805 (95% CI: 0.667–0.972, p = 0.024), close to the true value of 0.80.
| Method | Treatment HR | 95% CI | p-value |
|---|---|---|---|
| Naive (baseline) | NA | – | – |
| Time-dependent Cox | 0.805 | 0.667–0.972 | 0.024 |
The naive analysis cannot estimate the treatment HR because all patients start untreated. The time-dependent Cox correctly identifies HR = 0.805 (p = 0.024).
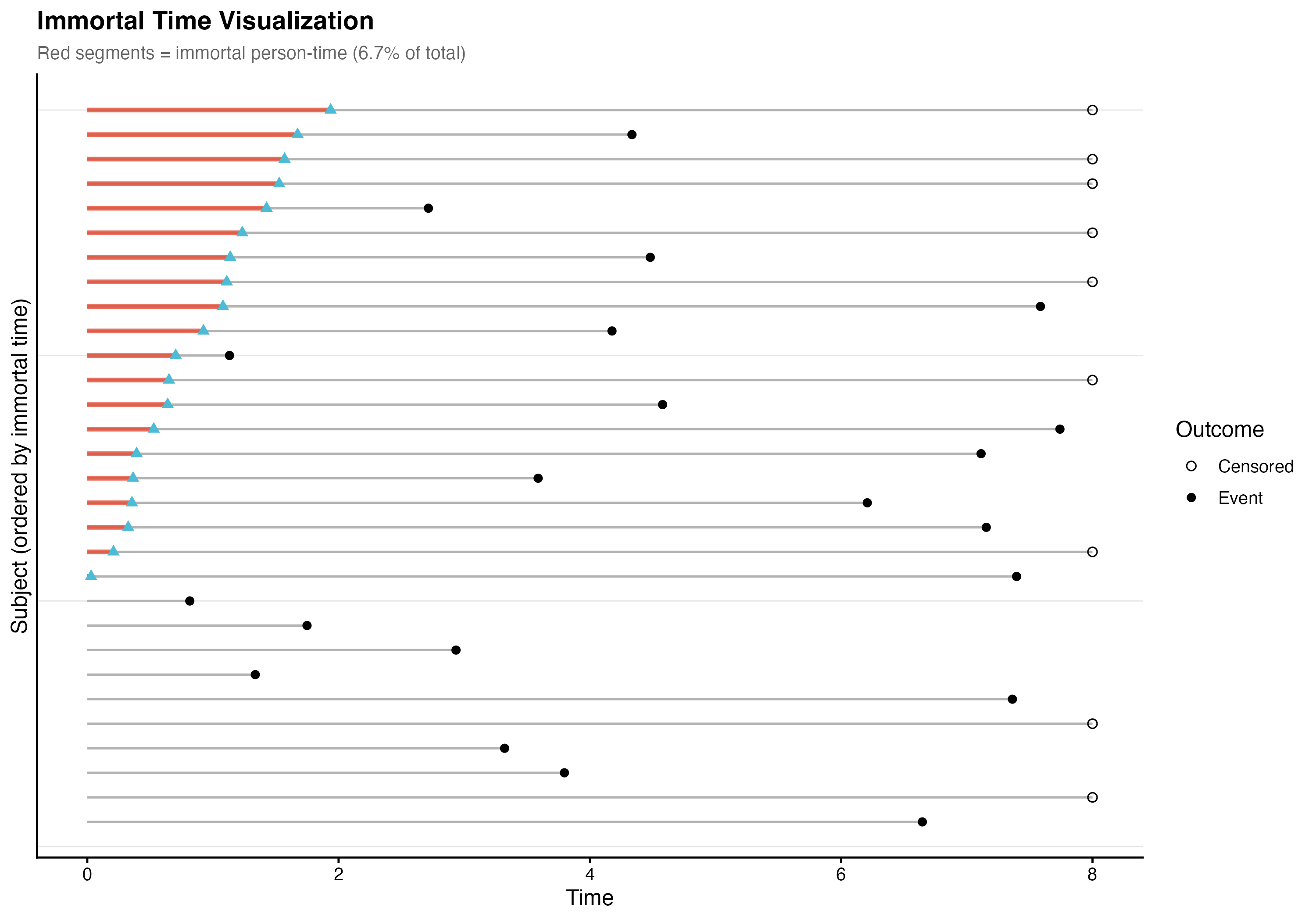
7.7.2 Immortal Time Quantification
Of the total 3736 person-time units of follow-up, 249.5 units (6.68%) constituted immortal person-time — time during which switchers were misclassified as treated but had not yet received treatment. The mean immortal time per switcher was 0.93 time units (median 0.83).
| Metric | Value |
|---|---|
| Total person-time | 3736.0 |
| Total immortal time | 249.5 |
| % immortal | 6.68% |
| Mean immortal time per switcher | 0.93 |
| Median immortal time per switcher | 0.83 |
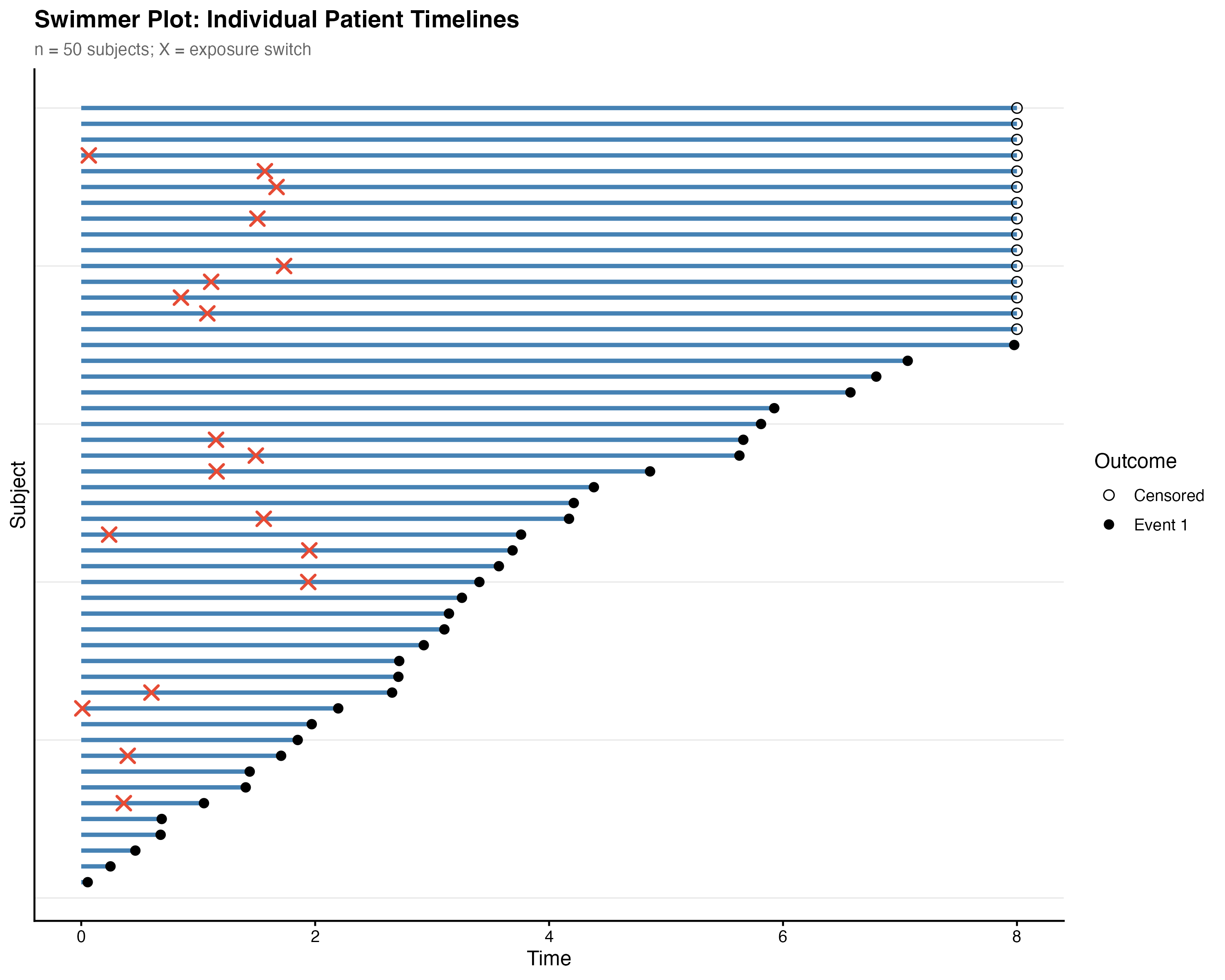
7.7.3 Swimmer Plot
7.7.4 Landmark Analysis
